Through selective breeding, gains have been made in both Terminal and Replacement traits in beef cattle in Ireland. This usually happens through the identification and proliferation of superior lines for traits of interest. Consequently, a smaller number of animals get selected which can lead to a reduction in genetic diversity in a population.
Genetic Diversity is defined as the variation in the amount of genetic information within and among individuals of a population, a species or a breed. For a population to survive it is important to reduce the loss of genetic variation in a population. It is very important to maintain genetic diversity for the future survival of the breed.
Inbreeding is defined as the probability that two alleles are identical by descent and occurs when related individuals are mated to each other (McParland, et al., 2006). The question then arises; what is the problem with inbreeding? Simply put, inbreeding results in a decline in performance of the resulting progeny. This is referred to as inbreeding depression. Inbreeding depression particularly affects less visible traits like reproductive and health traits, along with the more visible, growth, lactation and survival. Inbreeding depression was expressed as a reduction in post weaning gain of 240g per percentage increase in inbreeding in the US Limousin population (Gengler et al., 1998).
Inbreeding has been shown to negatively impact the live skeletal and muscling traits of Irish beef breeds (McParland, et al., 2008), particularly affecting loin development, hindquarter development, depth of rump and condition score. Some research has reported a reduction in scrotal circumference in yearling Hereford bulls (Smith, et al., 1989) which could potentially impact on male fertility. The higher the level of inbreeding in an individual the greater the inbreeding depression. Mating of related individuals may also lead to increased embryonic mortality and prevalence of genetic defects.
As selection intensity increases through reproductive methods like embryo transfer and artificial insemination, it can result in a fewer number of parents to provide the next generation of breeding animals. The possibility of selecting related animals is also enhanced by the methods animal breeders use to select animals, like breeding values e.g. €urostars. Increased usage of these techniques will in turn lead to an increased rate and level of inbreeding in the resulting population.

When looking at inbreeding within a population, it is important to look at the animals with a large genetic contribution to the population. This does not necessarily mean the bulls with the largest number of progeny, it is more to do with how many times they appear in the pedigrees of the reference population. For this analysis, the reference population is pedigree females, still alive, older than one year and born after the year 2000.
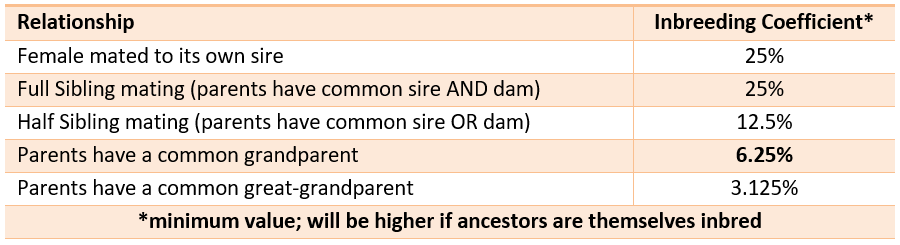
Many breeding animals are related to the bulls with large marginal contributions, so it is important to also look at the other measures around inbreeding. Many bulls have progeny and a small marginal contribution to the breed. You can use the relationship figures to assess what the inbreeding of an offspring would be. E.g., if a bull has an average relationship of 5% to the current pedigree population; this means, on average, the inbreeding percentage of a mating between the bull and the average dam would result in a progeny with an inbreeding level of 2.5%.
The analysis of inbreeding here has been carried out based on pedigree information in the ICBF database. Going forward, with more animals genotyped, there is an opportunity to carry out analysis using genomic inbreeding. Genomic inbreeding measures the relationship between two animals by assessing the level of homozygosity in their genes, and therefore providing a more accurate measure of inbreeding of an animal. We can identify animals that share the same genes, even though they may not be related through pedigree. This information is critically important when creating breeding strategies, to avoid the mating of animals that are carriers for genetic diseases. As breeders usually check pedigrees when planning matings, genomic inbreeding can provide a further insight into the genetic make-up of the animals, as while there may be an overlap in the pedigree of two animals, the reality may be that these animals have no genes in common. This helps to increase the number of potential sires a breeder could choose from when planning matings.
Going forward, advancements around genomic inbreeding will help breeders make more informed decisions when constructing breeding strategies.
This article has been prepared by ICBF in good faith on the basis of information provided to it. No representation or warranty expressed or implied is made or given by ICBF as to the accuracy, reliability, completeness of this article. ICBF shall not be liable for any losses (weather direct or indirect), damages, costs or expenses whatsoever, incurred or arising from any use of or reliance on this article or the information contained in it by any person.
